Reference: Development of a quantitative COVID-19 multiplex assay and its use for serological surveillance in a low SARS-CoV-2 incidence community
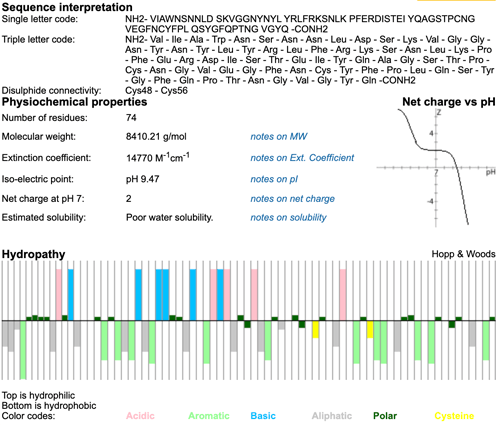
...A synthetic peptide of the RBD region of the Spike protein (synthetic RBD) was a commercially available (LifeTein LLC, Somerset, NJ, product number LT5587). This 74 amino acids peptide (aa 433 to 506, YP_009724390) was composed of the following sequence: NH2- VIAWNSNNLDSKVGGNYNYL YRLFRKSNLKPFERDISTEI YQAGSTPCNGVEGFNCYFPL QSYGFQPTNGVGYQ- CONH2, including a disulfide bridge: Cys480-Cys488...
Researchers from around the world have asked LifeTein for providing peptides to develop vaccines targeting the novel coronavirus (2019-nCoV) or SARS-CoV-2.
LifeTein synthesized the receptor-binding motif (RBM) of 2019 Coronavirus SARS-CoV-2 surface glycoprotein in 6 days. This receptor-binding domain (RBD) a critical determinant of virus-receptor interaction and thus of viral host range and tropism. The RBD also includes important viral-neutralizing epitopes (21–23), and it may be sufficient to raise a protective antibody response in inoculated animals.
A total of nine cysteine residues are found in the RBD, six of which forming three pairs of disulfide bonds. Among these three pairs, two are in the core (Cys336-Cys361 and Cys379-Cys432) to help stabilize the beta sheet structure while the remaining one (Cys480-Cys488) connects loops in the distal end of the RBM.
Studies showed that the sequence of 2019-nCoV coronavirus RBD, including its receptor -binding motif (RBM) that directly contacts ACE2 and uses ACE2 as its receptor with much higher affinity (10-20 times higher!) than SARS.
Catalog number: LT5587, 74aa
Peptide sequence: NH2- VIAWNSNNLD SKVGGNYNYL YRLFRKSNLK PFERDISTEI YQAGSTPCNG VEGFNCYFPL QSYGFQPTNG VGYQ -CONH2
Three letter sequence: NH2- Val - Ile - Ala - Trp - Asn - Ser - Asn - Asn - Leu - Asp - Ser - Lys - Val - Gly - Gly - Asn - Tyr - Asn - Tyr - Leu - Tyr - Arg - Leu - Phe - Arg - Lys - Ser - Asn - Leu - Lys - Pro - Phe - Glu - Arg - Asp - Ile - Ser - Thr - Glu - Ile - Tyr - Gln - Ala - Gly - Ser - Thr - Pro - Cys - Asn - Gly - Val - Glu - Gly - Phe - Asn - Cys - Tyr - Phe - Pro - Leu - Gln - Ser - Tyr - Gly - Phe - Gln - Pro - Thr - Asn - Gly - Val - Gly - Tyr - Gln -CONH2
Modifications:Disulfide Bridge: Cys480-Cys488
Quantity: 1mg
Purity: >97% by HPLC
Molecular Weight: 8410.21 g/mol
Download Data File: HPLC
HPLC  Mass
Mass