Disulfide-bridged peptides, head-to-tail cyclic peptides, side-chain cyclization, and advanced peptide macrocyclization services.
LifeTein provides custom cyclic peptide synthesis services for research applications requiring enhanced peptide stability, conformational constraint, receptor selectivity, or improved resistance to enzymatic degradation. Our team routinely prepares cyclic peptides through disulfide bond formation, amide bond cyclization, lactam bridge formation, and cysteine-selective macrocyclization strategies.
Cyclic peptides can be designed through several different linkage strategies depending on the sequence, functional groups, ring size, and desired biological application. Common cyclization formats include:
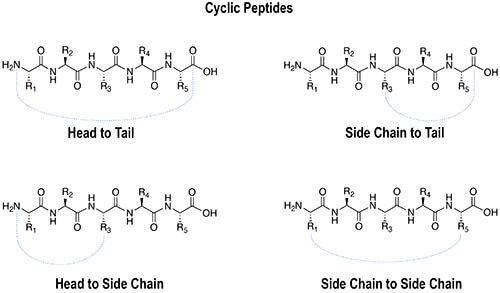
Disulfide bonds are important structural elements in many extracellular peptides and proteins. A disulfide bridge forms when the thiol groups of two cysteine residues are oxidized to create a covalent S–S bond. This linkage can stabilize peptide conformation, reduce flexibility, and help maintain biological activity.
For synthetic peptides, disulfide bond formation requires careful control of cysteine protection, deprotection, oxidation conditions, peptide concentration, pH, and reaction time. For peptides containing multiple cysteine residues, selective protection strategies are often required to guide correct disulfide pairing and reduce scrambling.
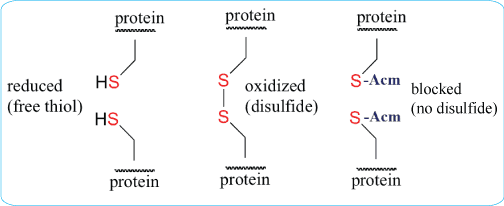
For simple peptides containing one cysteine pair, disulfide formation is often achieved by controlled oxidation of the purified linear peptide. For more complex peptides containing two or three disulfide bridges, orthogonal cysteine protecting groups may be used to direct stepwise disulfide formation.
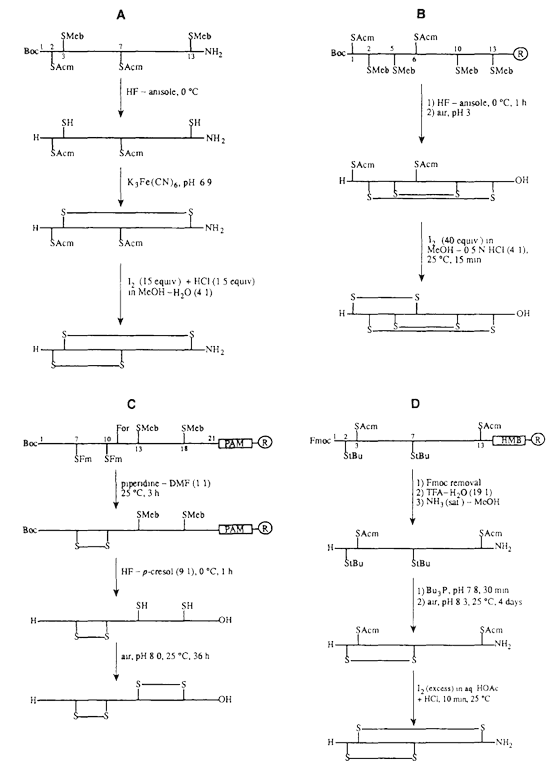
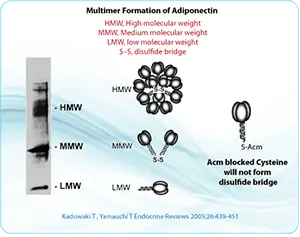
Peptides with multiple disulfide bonds are technically challenging because incorrect cysteine pairing can produce disulfide isomers, dimers, multimers, or misfolded products. LifeTein applies sequence-specific synthetic planning to improve the probability of obtaining the desired disulfide connectivity.
Key considerations include:
LifeTein has experience synthesizing cysteine-rich peptides such as defensin-like peptides containing three disulfide bonds. These peptides often require careful oxidation control because they are compact, cationic, and prone to aggregation or disulfide scrambling.
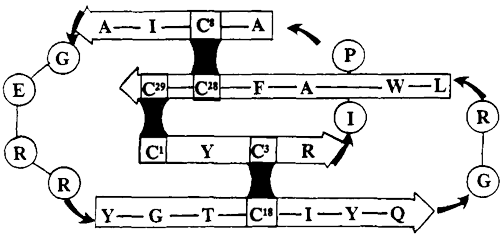
Oxidation conditions are selected based on peptide sequence and solubility. Typical parameters may include dilute peptide concentration, controlled pH, mild oxidizing reagents, and reaction monitoring by analytical HPLC. The optimal pH is commonly in the mildly acidic to neutral range, depending on peptide behavior.
Head-to-tail cyclic peptides are formed by creating an amide bond between the N-terminal amino group and C-terminal carboxyl group. This removes free termini and can significantly improve resistance to exopeptidase degradation.
Amide-based peptide cyclization can also be performed through side-chain functional groups. Common examples include lactam bridges formed between lysine and aspartic acid or lysine and glutamic acid residues.
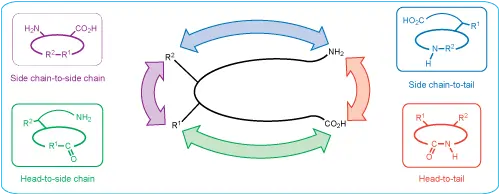
Successful amide cyclization depends strongly on ring size, steric hindrance, sequence flexibility, protecting group strategy, and dilution conditions. In some cases, pseudodilution or resin-assisted strategies may be used to favor intramolecular cyclization over dimerization or oligomerization.
For peptides with multiple cysteine residues, selective cysteine protection is often essential. Protecting groups such as Acm, Trt, tBu, Mob, Mmt, and related groups can be used to control when each cysteine becomes available for oxidation.
S-acetamidomethyl cysteine, Cys(Acm), is particularly useful because it is stable under many acidic and basic conditions and compatible with common peptide synthesis workflows. Acm protection can be used to block unwanted disulfide formation while preserving the cysteine residue in the peptide sequence.
This strategy is useful when:
In addition to classical disulfide and amide cyclization, LifeTein can support selected macrocyclization strategies designed to improve peptide rigidity, proteolytic stability, binding affinity, and cell permeability.
Perfluoroarene-based peptide stapling uses cysteine-selective chemistry to generate rigid perfluoroaromatic linkages. This approach can introduce a lipophilic, conformationally restrictive staple and may improve binding, stability, or membrane permeability depending on the peptide sequence and target.
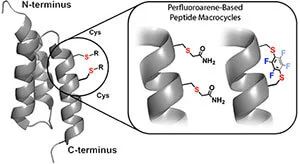
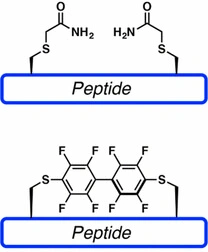
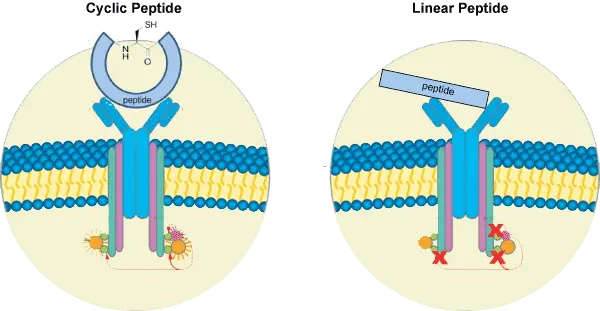
Compared with linear peptides, cyclic peptides often show improved metabolic stability, enhanced conformational control, and increased target selectivity. Because cyclic peptides lack free termini or contain a constrained structure, they are often less susceptible to degradation by exopeptidases.
| Linear Peptides | Cyclic Peptides |
|---|---|
| More conformational flexibility | Conformationally constrained structure |
| Often lower metabolic stability | Improved resistance to enzymatic degradation |
| Free N- and C-termini may be susceptible to exopeptidases | Reduced terminal degradation, especially in head-to-tail cyclic peptides |
| May show lower receptor selectivity | May improve binding specificity and potency |
RGD peptides are widely used in integrin-binding studies. Cyclization can restrict peptide conformation and improve stability compared with the corresponding linear sequence.
In one study comparing linear and cyclic RGD peptides, the cyclic peptide showed significantly improved solution stability at neutral pH. The enhanced stability was attributed to the constrained ring structure, which limited backbone degradation pathways associated with the Asp residue.
Reference: Solution stability of linear vs. cyclic RGD peptides. J Pept Res. 1999 May;53(5):530–541.
Publication example: A CPP-Ts peptide containing 45 amino acid residues and three properly positioned disulfide bonds, along with a 26-residue CPP-Ts subpeptide, was chemically synthesized by LifeTein, LLC.
View the Scientific Reports publication →
Please send us your peptide sequence, desired cyclization type, purity, quantity, and any special requirements. If the disulfide connectivity or cyclization site is known, please include that information in your quote request.
You may also download and complete the Peptide Quotation Form.
Tip: How to detect small peptides clearly and sensitively by Western blotting or SDS-PAGE?