Peptide libraries are widely used to study protein–peptide interactions, identify active binding regions, optimize peptide leads, and support epitope mapping, enzyme profiling, and related screening workflows. LifeTein provides peptide library synthesis services for research groups needing structured peptide sets for discovery, mapping, and assay development.
Peptide libraries can be prepared in multiple formats depending on the project, including overlapping peptide sets, truncation libraries, substitution libraries, random libraries, positional libraries, and array-based presentations for screening applications.
| Main uses | Epitope mapping, peptide scanning, interaction studies, screening, and peptide lead optimization |
| Formats | Individual vials, cellulose support, peptide arrays, discs, and other project-appropriate screening formats |
| Typical length | Commonly 3–30 amino acids; longer libraries may also be discussed depending on sequence and project design |
| Minimum library size | 20 peptides |
| Quality options | Crude and purified libraries with MS and optional HPLC support depending on project type |
Overlapping peptide libraries can be used to identify binding regions in proteins and are widely applied in antibody epitope mapping and protein–protein interaction studies.
Truncation, alanine scanning, and positional substitution libraries can help define the minimal active region and identify residues important for activity or binding.
Random and combinatorial peptide libraries can support the identification of active sequences when the preferred motif is not yet known.
Depending on the workflow, peptide sets may be delivered in individual vials, on cellulose support, as discs, or in array-compatible formats for screening use.
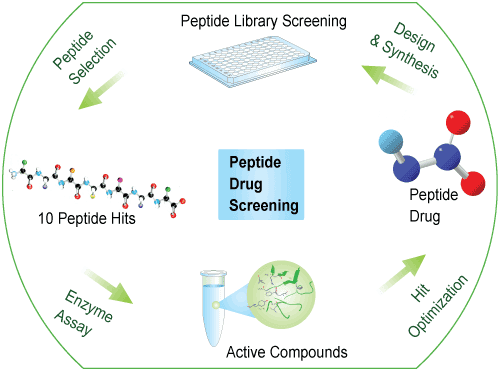
Overlapping peptide library
A protein sequence is divided into overlapping fragments. This format is commonly used for continuous epitope mapping and linear binding-region studies.
Truncation peptide library
A known peptide is systematically shortened to identify the minimum region needed for activity or binding.
Alanine scanning library
Each residue is systematically replaced by alanine to identify positions important for peptide conformation, activity, or target recognition.
Random peptide library
Selected positions are substituted broadly to explore sequence space and identify improved activity or altered specificity.
Scrambled peptide library
Sequence variants based on rearrangement of the original peptide are often used as controls or as part of screening workflows to test sequence dependence.
Positional peptide library
Specific regions or positions within a peptide are replaced systematically to evaluate how sequence variation affects function.
Cellulose-bound peptide arrays remain an important research format for peptide screening and interaction studies. These arrays can support large numbers of peptide spots on a single membrane and are useful in repeated probing workflows when the chemistry and regeneration conditions are appropriate.
Reference: Cellulose-bound Peptide Arrays: Preparation and Applications, Biotechnology and Genetic Engineering Reviews, Vol. 24, 31–106 (2007).
Different support/linker systems may be used depending on whether the array is intended for single use, multiple use, or specific assay conditions. In peptide-array projects, design of linker chemistry, peptide orientation, and display format is often as important as the sequence set itself.
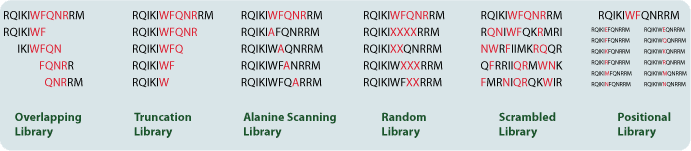
Synthetic peptide libraries are also valuable in screening for enzyme substrates, inhibitors, cell-binding peptides, and protein-domain ligands. Because many molecules can be handled in parallel, peptide library approaches are useful in discovery settings where systematic sequence exploration is needed.
Example from the literature
Overlapping peptides from LifeTein were used for epitope mapping in: Cell, Volume 159, Issue 4, 6 November 2014, Pages 844–856, DOI: 10.1016/j.cell.2014.10.032
Another example cited LifeTein pools of overlapping 15-mer peptides in a peptide-library immunology workflow: Phase 2 trial of T-cell activation using MVI-816 and pembrolizumab in patients with metastatic, castration-resistant prostate cancer (mCRPC)
LifeTein’s peptide library program can synthesize libraries in individually labeled vials or other appropriate delivery formats depending on the application. Each peptide undergoes quality control appropriate to the selected synthesis and purification format.
Pricing depends on the number of peptides, peptide length, purity target, modification requirements, and final delivery format. For practical quoting, peptide library projects are best evaluated based on the exact design rather than using a single universal library price table.
| Library type | Typical considerations | Quotation |
|---|---|---|
| As synthesized / crude libraries | Sequence-dependent crude purity, scale, and delivery format | Request Quote |
| Purified libraries | Target purity, number of peptides, analytical requirements, and modifications | Request Quote |
| Array or membrane-based projects | Array format, surface chemistry, peptide orientation, and assay workflow | Request Quote |
How to start
Send us your sequence set, parent protein sequence, scan design, library type, desired purity, and preferred delivery format. If you already have a quote number, you may also proceed directly to place an order.
Please email your inquiry to sales@lifetein.com or use our online quotation form. We can help define the most practical library design and delivery format for your project.