Custom biotin-labeled peptides for pull-down assays, ELISA, peptide-protein interaction studies, affinity capture, screening assays, and streptavidin-based detection.
LifeTein provides custom biotinylated peptide synthesis with flexible labeling positions, spacer options, and matched control peptides. Biotinylated peptides are widely used as affinity probes because the biotin–streptavidin interaction is exceptionally strong and highly useful for immobilization, capture, detection, and enrichment workflows.
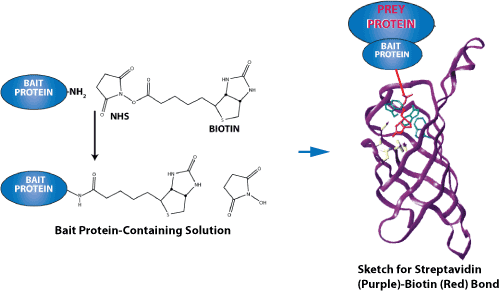
Biotinylated peptides can be immobilized on streptavidin-coated plates, membranes, slides, or beads. The peptide serves as bait, while interacting proteins, antibodies, or other binding partners are captured and analyzed.
A biotinylated peptide can be used as bait to identify peptide-binding proteins. In a typical pull-down workflow, the biotin-labeled peptide is immobilized on streptavidin or avidin beads and incubated with a biological sample such as cell lysate, nuclear extract, serum, or purified recombinant protein.
After incubation, unbound proteins are removed by washing. Bound proteins can then be eluted and analyzed by SDS-PAGE, western blot, mass spectrometry, or other downstream methods. Comparing binding to a modified peptide and a matched control peptide helps identify sequence-specific or modification-dependent interactions.
Biotinylated peptides are especially useful for peptide ELISA because they can be immobilized directionally on streptavidin-coated 96-well plates. This format provides a convenient way to screen antibodies, test binding activity, compare peptide variants, or validate phospho-specific and modification-specific antibodies.
For antibody development projects, LifeTein can synthesize both the modified peptide antigen and matched control peptides. These peptide pairs are useful for ELISA screening, dot blot validation, peptide competition assays, and affinity purification.
The position and spacer length of the biotin label can strongly affect assay performance. Direct biotin attachment may work for many short peptides, but a spacer is often recommended when the peptide needs to remain accessible after immobilization.
| Format | Best Use | Key Advantage |
|---|---|---|
| Biotin-peptide | Basic immobilization and detection | Simple, economical labeling |
| Biotin-Ahx-peptide | ELISA, pull-down, antibody screening | Improves peptide accessibility |
| Biotin-PEG-peptide | Hydrophobic or sterically restricted peptides | Improves flexibility and spacing from the surface |
If you are unsure which design is best, LifeTein can help select the appropriate biotin position and spacer based on your assay format.
The biotin-streptavidin interaction is one of the strongest non-covalent biological interactions and is widely used in biochemical assays. Streptavidin is tetrameric and can bind up to four biotin molecules, making it useful for surface immobilization, bead capture, and signal amplification.
Because this interaction is extremely stable, standard biotinylated peptides are ideal when strong capture is desired. If downstream release of the captured biomolecule is needed, consider cleavable or photocleavable biotin designs.
Learn more about photocleavable biotin peptides →
Biotin-based labeling and capture systems continue to support important studies in immunology, cell-cell interaction mapping, and peptide/protein interaction analysis.
Please send us your peptide sequence, desired biotin position, spacer preference, purity, quantity, and intended assay application. If you are designing a pull-down or ELISA experiment, please also indicate whether you need matched control peptides.