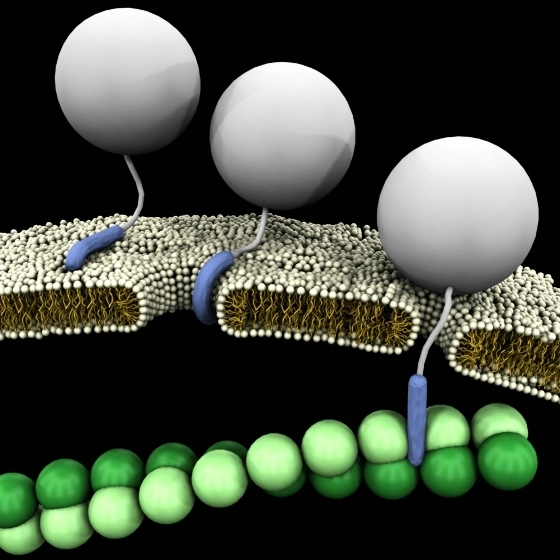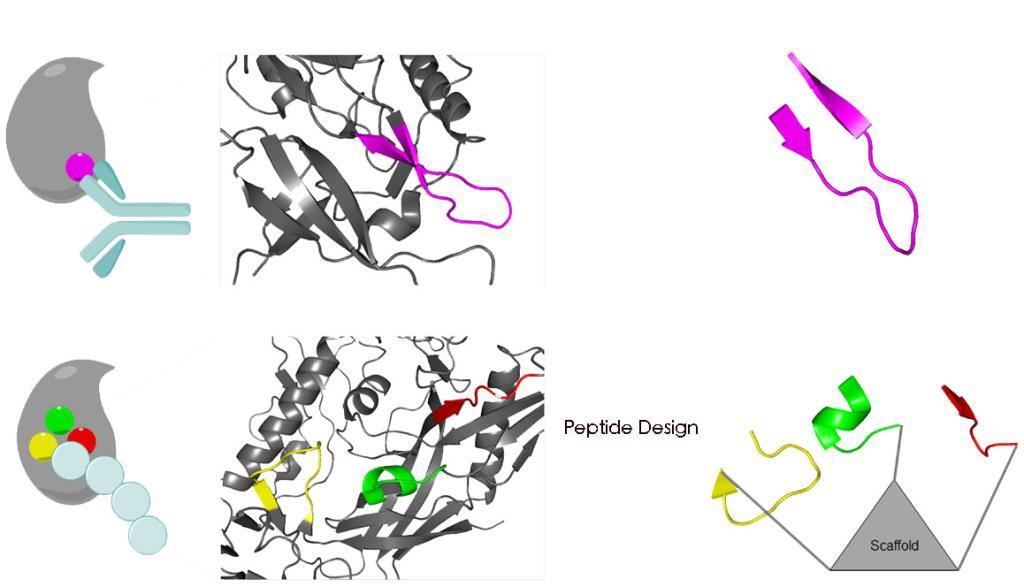
Synthetic peptides have proven an excellent type of molecule for the mimicry of protein sites. The modified peptides increase the proteolytic stability of the molecules, enhancing their utility for biological applications.
Toolbox for Peptide Synthesis: Non-Proteinogenic Amino Acids and Site-Selective Ligation
The long peptides can be synthesized by the ligation method. Amino acid derivatives with modified backbone length and side-chain orientation, such as d-amino acids, N-alkyl glycine monomers, or proteolytically stable amino acid derivatives can be introduced to the peptides.
Protein Secondary Structure Mimics: α-Helix Mimics, β-Sheet Mimics
Peptide chains can be organized into secondary structures, such as α-helices and β-sheets. Peptides that mimic α-helices and β-sheets of proteins are attractive targets for drug development and tools to explore protein binding mechanism.
The α-helical conformation of a peptide can be induced by adding covalent links between amino acid side chains at selected positions. These links can be formed by lactam and disulfide bridges, triazole-based linkages, and hydrocarbon staples.
In β-sheets, β-strands are connected via loops or turns. Methods to mimic turn structures include macrocyclization, dipeptide of d-proline and l-proline, or α-aminoisobutyric acid in combination with either a d-α-amino acid or an achiral α-amino acid. An example of stimuli-responsive peptides is the temperature-dependent formation of hydrogels by β-sheet peptides. The β-hairpin mimic undergoes gelation upon heating at 60°C, and is completely reversible while cooling.
Protein Mimics in Biomedical Research
Peptides mimicking the CHR region of gp41 were developed to inhibit the formation of the six-helical bundle. Peptides that mimic these receptors are useful tools to explore the details of virus infection mechanism, as well as to develop new drugs against HIV-1. Peptides that mimic the extracellular domains of seven transmembrane G protein-coupled receptors (GPCRs), which is composed of the N-terminus (NT) and the three extracellular loops (ECLs) were explored. Peptide Ac-RERF-NH2 has a high propensity to adopt an α-turn structure and could be a promising drug candidate against cancer.
The design of peptides as protein mimics has evolved as a promising strategy for the exploration of protein-protein interactions, as they are biocompatible, biodegradable, and functionally selective.